Western blot sample formulations, including lysis buffers, lysate from cell culture, lysate from tissues and resolution of protein concentration.
Print this protocol.
Western blot protocol contents
- Lysis buffers
- Protease and phosphatase inhibitors
- Preparation of lysate from cell culture
- Preparation of lysate from tissues
- Resolution of protein concentration
- Preparation of samples for loading into gels: denatured and native, reduced and non-reduced
Lysis buffers
Lysis buffers differ within their capability to solubilize proteins, with individuals that contains sodium dodecyl sulfate (SDS) along with other ionic detergents regarded as the harshest and for that reason probably to own greatest yield.
The primary consideration when selecting a lysis buffer is whether or not the selected antibody will recognize denatured samples. When this isn't the situation, it will likely be noted around the antibody datasheet, and buffers without detergent or with relatively mild non-ionic detergents (NP-40, Triton X-100) ought to be used.
Protein location and lysis buffer choice
RIPA or use mitochondrial fraction protocol*
*Proteins which are found solely or predominantly inside a sub-cellular location could be more filled with a lysate from the sub-cellular fraction in contrast to whole cell or tissue lysates. This is often helpful when attempting to acquire a signal for any weakly-expressed protein. Please consult our separate protocols for sub-cellular fractionation.
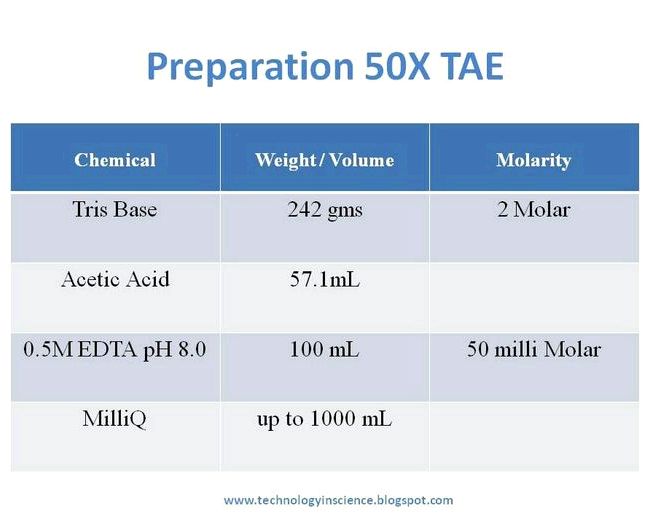
Lysis buffer recipes:
- 150 mM sodium chloride
- 1.% NP-40 (Triton X-100 could be substituted with NP-40)
- 50 mM Tris pH 8.
This can be a popular buffer for studying proteins which are cytoplasmic or membrane-bound, or whole cell extracts. If there's concern the protein of great interest isn't being completely obtained from insoluble material or aggregates, RIPA buffer might be more appropriate because it contains ionic detergents which will more readily bring the proteins into solution.
RIPA buffer (radioimmunoprecipitation assay buffer)
- 150 mM sodium chloride
- 1.% NP-40 or Triton X-100
- .5% sodium deoxycholate*
- .1% SDS (sodium dodecyl sulfate)
- 50 mM Tris, pH 8.
*Could be prepared like a 10% stock solution, which should be protected against light.
RIPA buffer is helpful for whole cell extracts and membrane-bound proteins, and could be more suitable to NP-40 or Triton X-100-only buffers for removing nuclear proteins. It'll disrupt protein-protein interactions and could therefore be problematic for immunoprecipitations and pull-lower assays.
In instances where you should preserve protein-protein interactions in order to minimize denaturation, a buffer without ionic detergents (eg SDS) and ideally without non-ionic detergents (eg Triton X-100) ought to be used.
Cell lysis with detergent-free buffer is achieved by mechanical shearing, frequently having a Dounce homogenizer or by passing cells via a syringe tip. In these instances, an easy Tris buffer will suffice, but because noted above, buffers with detergents are needed to produce membrane- or cytoskeleton-bound proteins.
- 20 mM Tris-HCl, pH 7.5
Tris-Triton buffer (cytoskeletal proteins)
- 10 mM Tris, pH 7.4
- 100 mM NaCl
- 1 mM EDTA
- 1 mM EGTA
- 1% Triton X-100
- 10% glycerol
- .1% SDS
- .5% deoxycholate
All of those buffers could keep at 4°C for many days or for approximately annually if split into aliquots and stored at -20°C.
Protease and phosphatase inhibitors
When lysis occurs, proteolysis, dephosphorylation and denaturation begin. These occasions could be slowed lower considerably if samples are stored on ice or at 4°C whatsoever occasions and appropriate inhibitors are added fresh towards the lysis buffer.
Ready-to-use cocktails of inhibitors from various suppliers can be found but you may make your personal cocktail.
Dilute in water. Don't re-use once defrosted.
Sodium orthovanadate preparation
Perform all stages in a fume hood.
- Make a 100 mM solution in double sterilized water.
- Set pH to 9. with HCl.
- Boil until without color. Minimize volume change because of evaporation by covering loosely.
- Awesome to 70 degrees.
- Set pH to 9. again.
- Boil until without color.
- Continue doing this cycle before the solution remains at pH 9. after boiling and cooling.
- Bring to the initial volume with water.
- Store in aliquots at -20°C. Discard if samples turn yellow.
Preparation of lysate from cell culture
- Put the cell culture dish on ice and wash cells with ice-cold PBS.
- Aspirate the PBS, adding ice-cold lysis buffer (1 mL per 10 7 cells/100 mm dish/150 cm 2 flask .5 mL per 5x10 6 cells/60 mm dish/75 cm 2 flask).
- Scrape adherent cells from the dish utilizing a cold plastic cell scraper, then lightly transfer the cell suspension right into a pre-cooled microcentrifuge tube. Alternatively cells could be trypsinized and washed with PBS just before resuspension in lysis buffer inside a microcentrifuge tube.
- Maintain constant agitation for 30 min at 4°C.
- Centrifuge inside a microcentrifuge at 4°C. You might want to vary the centrifugation pressure and time with respect to the cell type a guideline is 20 min at 12,000 revoltions per minute however this should be determined for the experiment (e.g. leukocytes require a very light centrifugation).
- Lightly take away the tubes in the centrifuge and put on ice, aspirate the supernatant and put inside a fresh tube stored on ice, and discard the pellet.
Preparation of lysate from tissues
- Dissect the tissue of great interest with clean tools, on ice preferably, so that as rapidly as you possibly can to avoid degradation by proteases.
- Put the tissue in round-bottom microcentrifuge tubes or Eppendorf tubes and immerse in liquid nitrogen to snap freeze. Store samples at -80°C for later or continue ice for fast homogenization.
- For any
5 mg bit of tissue, add
300 L of cold lysis buffer quickly towards the tube, homogenize by having an electric homogenizer, rinse the blade two times with another 2 x 300 L lysis buffer, then maintain constant agitation for just two h at 4°C (eg put on an orbital shaker within the fridge). Volumes of lysis buffer should be determined with regards to the quantity of tissue present. Protein extract shouldn't be too diluted to prevent lack of protein and enormous volumes of samples to become loaded onto gels. The minimum suggested concentration is .1 mg/mL, optimal concentration is 1–5 mg/mL).
Resolution of protein concentration
- Execute a Bradford assay, a Lowry assay or perhaps a bicinchoninic acidity (BCA) assay. Bovine serum albumin (BSA) is really a commonly used protein standard.
- After you have determined the power of each sample, you are able to freeze them at -20°C or -80°C for later or get ready for immunoprecipitation or loading onto a gel.
Preparation of samples for loading into gels
Antibodies typically recognize a little area of the protein of great interest (known as the epitope) which domain may reside inside the 3D conformation from the protein. To allow access from the antibody for this portion it's important to unfold the protein, ie denature it.
To denature, make use of a loading buffer using the anionic detergent sodium dodecyl sulfate (SDS), and boil the mix at 95–100°C for five min. Heating at 70°C for 5–10 min can also be acceptable and could be more suitable when studying multi-pass membrane proteins. These have a tendency to aggregate when steamed and also the aggregates might not go into the gel efficiently.
The conventional loading buffer is known as 2X Laemmli buffer (Laemmli United kingdom, 1970. Cleavage of structural proteins throughout the set up from the mind of bateriophage T4. Nature, 227, 680–5). It is also made at 4X and 6X strength to reduce dilution from the samples. The 2X will be included 1:1 ratio using the sample.
2x Laemmli buffer recipe
- 4% SDS
- 10% 2-mercaptoethanol
- 20% glycerol
- .004% bromophenol blue
- .125 M Tris HCl
- Look into the pH and produce it to pH 6.8
When SDS can be used with proteins, all the proteins become negatively billed by their attachment towards the SDS anions. SDS binds to proteins fairly particularly inside a mass ratio of just one.4:1. By doing this, SDS confers an adverse charge towards the polypeptide compared to the length. Denatured polypeptides become rods of negative charge with equal charge densities per unit length. Therefore, migration is dependent upon molecular weight, instead of the intrinsic control of the polypeptide.
SDS grade is essential for top-quality protein separation: a protein stained background along individual gel tracts with indistinct or slightly distinct protein bands are suggestive of old or low quality SDS. Inclusion of two-mercaptoethanol or dithiothreitol within the buffer reduces disulphide bridges, that is essential for separation by size.
Glycerol is put into the loading buffer to improve the density from the sample to become loaded and therefore keep up with the sample at the end from the well, restricting overflow and uneven gel loading.
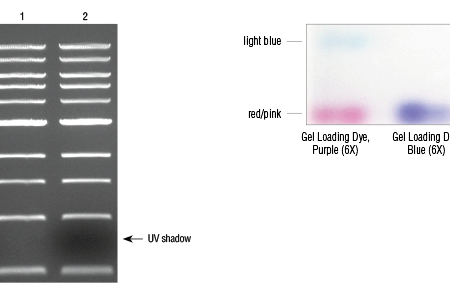
To visualise the migration of proteins it's quite common to incorporate a little anionic dye molecule within the loading buffer (eg bromophenol blue). Because the dye is anionic and small, it'll migrate the quickest associated with a component within the mixture to become separated and supply a migration front to watch the separation progress.
During protein sample treatment the sample ought to be mixed by vortexing pre and post the heating step for the best resolution.
Native and non-reduced samples
Alternatively, an antibody may recognize an epitope comprised of non-contiguous proteins. Even though the proteins from the epitope are separated from each other however sequence, they're near to one another within the folded three-dimensional structure from the protein, and also the antibody is only going to recognize the epitope because it exists on the top of folded structure.
During these conditions, you should operate a western blot in non-denaturing conditions, and will also be noted around the datasheet within the applications section. Generally, a non-denaturing condition means departing SDS from the sample and migration buffers and never heating the samples.
Certain antibodies only recognize protein in the non-reduced form (particularly on cysteine residues) and also the reducing agents -mercaptoethanol and DTT should be excluded from the loading buffer and migration buffer.